Hybridoma Sequencing Resources
When it comes to hybridoma sequencing, it can be confusing to determine which technology is appropriate for recovering antibody sequences. This blog will explain the main differences between the two main approaches to hybridoma sequencing; next-generation sequencing (NGS) and Sanger sequencing. These approaches have distinct features in terms of sequencing length, cost, volume, and diversity in antibody repertoire.
Sanger sequencing:
Before high-throughput sequencing, Sanger sequencing was the standard method of hybridoma sequencing. Sanger sequencing can sequence up to 96 samples with a turnaround time of ~5 days going from RNA extraction to determining the sequences of the variable region [1]. Sanger sequence is very cost-effective for a small number of samples. This method is adequate for identifying dominant clones or validating design objectives. However, a major limitation is the inability to sequence samples expressing multiple antibody sequences, also called polyclonal antibodies. In a recent study [2], ⅓ of hybridomas were found to be polyclonal. It is well known that the myeloma cell line used for fusions can also express its own antibody unrelated to the antibody of interest. These extra antibody transcripts make Sanger sequencing difficult on many hybridoma samples [3]. It would take an extra week to deconvolute the source DNA to monoclonal DNA through sub-cloning before resequencing. Overall, Sanger sequencing can be used for identifying monoclonal antibody sequences but the technology reaches its limit when researchers unknowingly have a hybridoma expressing multiple antibodies, and also need to scale up screening to the plate level.
[Next-Generation Sequencing (NGS):
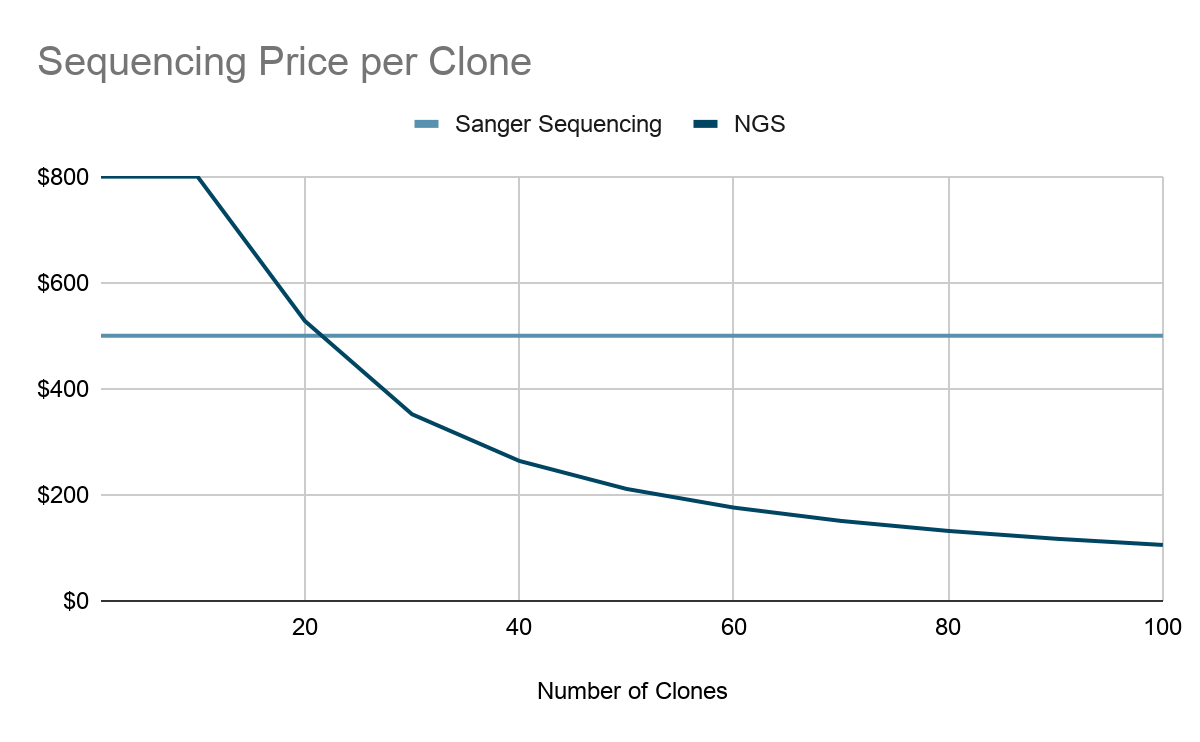
Next-Generation Sequencing is the new standard approach to hybridoma sequencing. It can cost ~$800 to sequence a clone [1]. The cost can go as low as $100 per hybridoma for sequencing at a high throughput scale. The biggest driver of cost savings with NGS is the ability to sequence many hybridomas in parallel in a single sequencing run. NGS can clearly provide more sequences per run which means low frequency variants in polyclonal samples can also be detected [2]. Overall, NGS has a great capability for high throughput screening, well suited for identifying monoclonal and polyclonal antibodies in a clone, and its sequencing depth can capture a more diverse antibody repertoire that Sanger wouldn’t be able to do.
References:
- Meyer L, López T, Espinosa R, Arias CF, Vollmers C, DuBois RM. A simplified workflow for monoclonal antibody sequencing. PLoS One. 2019;14(6):e0218717. Published 2019 Jun 24. doi:10.1371/journal.pone.0218717. (PubMed)
- Bradbury ARM, Trinklein ND, Thie H, Wilkinson IC, Tandon AK, Anderson S, Bladen CL, Jones B, Aldred SF, Bestagno M, Burrone O, Maynard J, Ferrara F, Trimmer JS, Görnemann J, Glanville J, Wolf P, Frenzel A, Wong J, Koh XY, Eng HY, Lane D, Lefranc MP, Clark M, Dübel S. When monoclonal antibodies are not monospecific: Hybridomas frequently express additional functional variable regions. MAbs. 2018 May/Jun;10(4):539-546. doi: 10.1080/19420862.2018.1445456. Epub 2018 Mar 29. PMID: 29485921; PMCID: PMC5973764. (Europe PMC)
- Rouet R, Jackson KJL, Langley DB and Christ D (2018) Next-Generation Sequencing of Antibody Display Repertoires. Front. Immunol. 9:118. doi: 10.3389/fimmu.2018.00118. (PubMed)
Relevant Articles:
Get In Touch
Tell Us About Your Project.
Need more information? We’re here to answer any questions you have.
Antibody Resources
Antibodies as Tools*
Antibody Production*
Antibody Reference Guide (downloadable)
Hybridoma Resources
Hybridoma Technology
Hybridoma Sequencing
Sequencing Pitfalls
Case Studies (Downloadable)
Reptor
Alicanto
Valens
*Stay tuned for these upcoming topics!
