Constructing an Accurate Antibody Repertoire
What is an antibody repertoire?
The immune repertoire is the collection of unique immunoglobulin and T-cell receptor sequences present in an individual at a particular time. At Digital Proteomics we focus primarily on the immunoglobulin, or B-cell receptor repertoires of humans and other mammals. The total number of B cells and plasma cells that encode full-length immunoglobulins in an adult human is estimated to be 10¹º-10¹¹, which puts an upper bound on the total size of the B-cell receptor, or antibody, repertoire. The antibody repertoire is dynamic, with sequence diversity and composition of the repertoire changing dramatically over time. In this post, we discuss the approach to immune repertoire construction that we employ in our Reptor™ and Alicanto® services. Before repertoire construction and analysis can begin, the B-cell receptor transcripts must be sequenced. For a bit about that process, read our previous post on immunosequencing.
Due to high sequence diversity, antibody repertoire sequencing and assembly of an individual repertoire is complex and requires specialized tools. It’s important to recognize the unique challenges of antibody repertoire sequence and analysis, which make them ill-suited for standard RNA-seq or target-enrichment workflows.
Antibody repertoire construction overview
Read quality filtering
As in any bioinformatic pipeline, for immune repertoire analysis ‘garbage in = garbage out’. Read filtering is an important part of the process, and the best practices for any RNA/DNA sequencing analysis apply. This includes removing reads with low quality, removing reads that are generated from a spiked-in library (such as PhiX), and trimming primer and sequencing adapter sequences. Looking at overall statistics for your run using a free tool such as FastQC is also a great way to spot problems in your sequencing run.
Tell Us About Your Project
Read stitching
In order to recover the full 330+ nucleotides of the variable region, paired-end reads or long single-end reads are needed. In our workflows, we use the Illumina MiSeq system to generate 2×300 nucleotide paired reads. This results in considerable overlap between reads. We construct a stitched read by identifying the overlap and constructing a consensus sequence in the overlapping region.
Length and quality filtering
Once read pairs have been stitched into a single consensus read, additional filtering is applied. Specifically, we remove consensus reads that are too short or too long, which can be signs that the read is truncated or was unstitchable. In our Reptor™ and Alicanto® workflows, we also apply a crude test of ‘antibody-ness’. The reads are the product of several rounds of PCR with antibody locus-specific primers, however, a subset of sequences may be to off-target loci. We quickly test whether each sequence roughly aligns to a reference germline V gene. If a decent alignment can’t be found between each read and a reference gene, the read is probably not an antibody (or the reference is woefully incomplete). At this stage, we collapse identical reads into a single read.
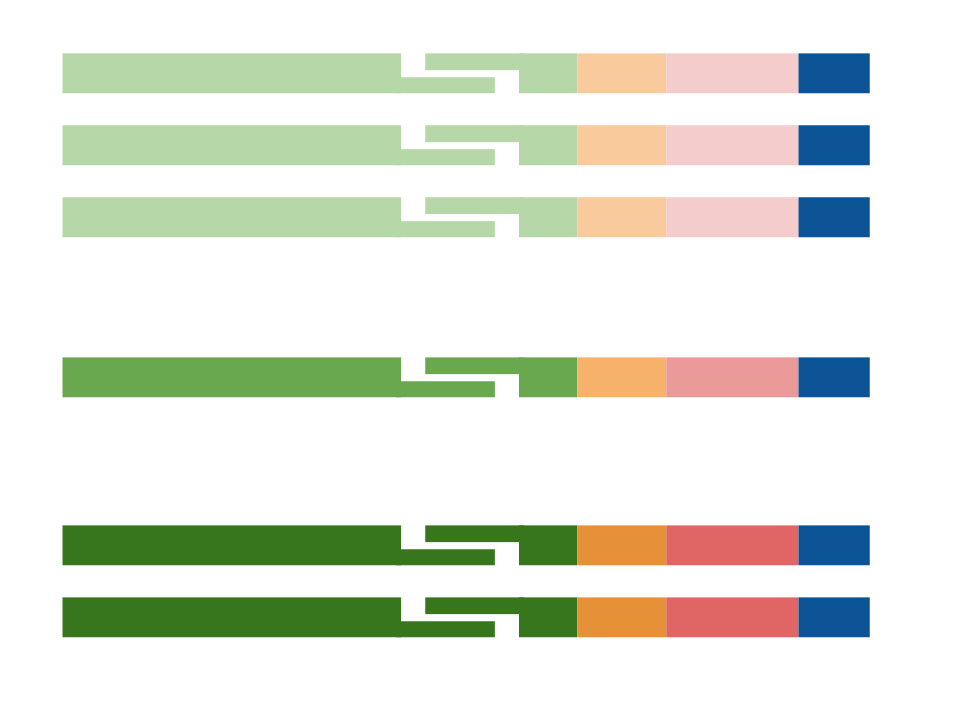
Error correction
A single amino acid change can dramatically alter the stability and affinity of an antibody. For that reason, constructing an error-corrected repertoire is a crucial first step to many downstream applications from antibody discovery to clone tracking. Both PCR amplification and sequencing can introduce errors, which must be corrected. A simple approach is to apply an abundance threshold and remove any sequences with fewer than a fixed number of reads. The idea behind this approach is that errors should be rare, and true antibody sequences should appear multiple times. Low abundance correct sequences will be dropped, removing real and important information about the repertoire.
Unique molecular identifiers (UMIs) have grown in popularity for correcting errors as well as accurately quantifying RNA abundance of each antibody locus (Khan et al. 2016, Turchaninova et al. 2016). At Digital Proteomics, our approach is based on the Hamming graph clustering approach to performing error correction developed at UCSD (Safonova et al. 2015). We create a node in the graph for each distinct antibody sequence, and create an edge between nodes if the sequences are within a predefined Hamming distance. In principle, erroneous sequences will appear in subgraphs also containing the correct sequence. A consensus sequence is created by collapsing dense subgraphs.
V(D)J labeling and complementarity-determining region identification
After error correction, the repertoire consists of a collection of corrected sequences. In order to further characterize the repertoire we must determine the germline sequences that gave rise to each antibody. This process is called V(D)J labeling, and open source tools such as IMGT/HighV-Quest, IgBLAST, MiXCR have been developed for this purpose. We apply the colored antibody graph technique (Bonissone and Pevzner, 2016) to determine the germline V, D, and J genes for each antibody sequence. In addition to identifying the original germ line sequence, aligning each antibody to the germline allows us to determine hypermutation sites.
Germline gene labeling also aides complementarity-determining region (CDR) identification. Each antibody chain has 3 CDRs, two of which reside fully in the V gene segment and a third which occurs at the junction of the V, D, and J (or V and J in the case of the light chains). Identifying these regions is important for understanding the relationships between antibodies in the repertoire, and their evolution in response to immunological challenge. In our process, we cluster nearly identical CDR3 sequences into a single group, or clone. The figure below demonstrates the process of V(D)J labeling, CDR3 identification, and CDR3 clustering.
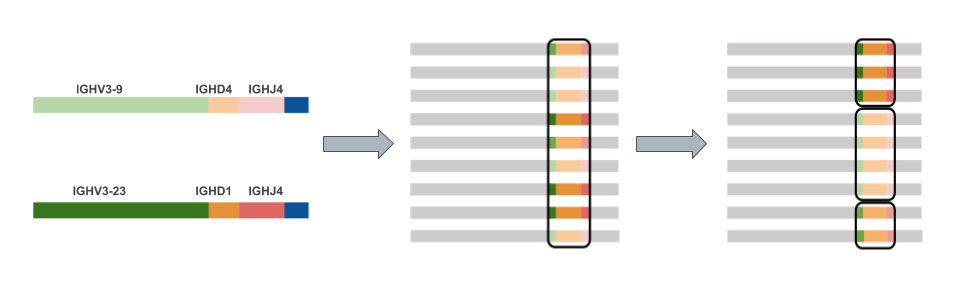
What was once tens of gigabytes of unintelligible reads is now an annotated antibody repertoire. The world is your oyster for antibody repertoire analysis, and we will survey that world in a future post. Stay tuned!
Tell Us About Your Project.
Need more information? We're here to answer any questions you have.
Resources
The Adaptive Immune Receptor Repertoire (AIRR) Community holds regular meetings and publishes standards for repertoire sequencing and analysis.
Practical guidelines for B-cell receptor repertoire sequencing analysis.
